SNIDDY Seminar Series: Ants, Bees, Penguins and Foxes
-
 Julien Riou
Julien Riou - 12 May, 2026

Our next online seminar will feature Nicolas Banholzer from the University of Copenhagen on Friday, March 13th 2026, 1–2pm
Title: Ants, Bees, Penguins and Foxes – Roles in the Socio-Molecular Transmission Network of the Danish Population during the SARS-CoV-2 Pandemic
Abstract: Two questions recurred during the COVID-19 pandemic: could targeted measures replace blanket lockdowns, and who should be vaccinated first? Both hinge on identifying which population subgroups contribute disproportionately to transmission. Answering this at population scale requires reconstructing who infected whom and linking those individuals to detailed sociodemographic information, which is rarely possible in most countries.
Denmark is a notable exception. In 2021, more than 290,000 SARS-CoV-2 genomes were sequenced from PCR-positive samples, with coverage consistently above 60%. The genomes can be linked through the Danish National Registers to individual-level data such as age and occupation, as well as to verified social relations like shared household, school, or workplace. From these data we build a “socio-molecular transmission network” consisting of plausible infector-infectee pairs with identical consensus sequences, with socially-related pairs prioritised over genome-only matches.
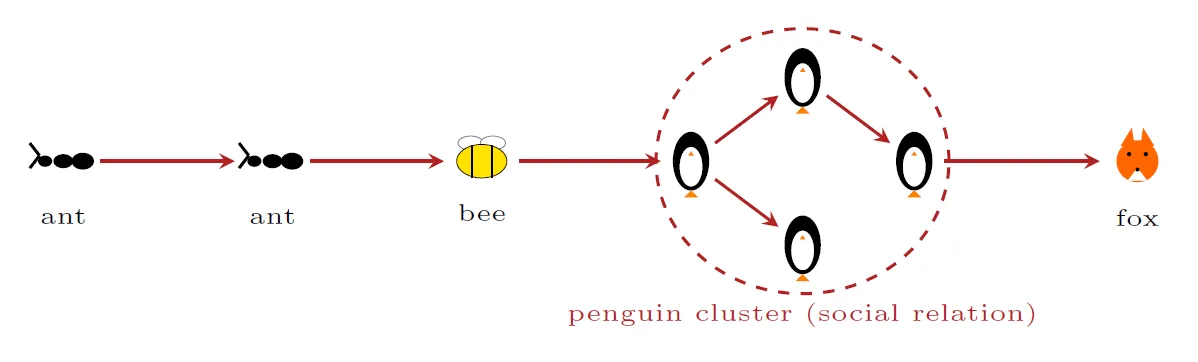
To identify influential subgroups, we compute individual-level network metrics and aggregate them by age group. We then assess each aggregated value against two complementary permutation-based null models. The first, shuffling, randomly permutes age-group labels across nodes while keeping the network topology fixed; the deviation from this null tells us whether a group’s individuals occupy structurally more (or less) influential positions than random individuals would. The second, shuffling and removal, additionally removes the target group from the network and recomputes the metric on the remaining nodes; comparing this drop to the drop expected when removing a random subset of equal size tells us how much the rest of the network’s transmission depends on that group’s presence. We apply this framework first to a metric that counts how many distinct individuals each node reaches within three transmission generations, and second to a categorisation of four pairwise transmission roles: ants connecting to other ants on a chain to a bee; bees pollinating a “penguin” cluster of socially-related individuals; penguins themselves; and foxes who infect no one onwards. The talk presents this framework and intermediate results from our exploratory safari, discussing what the role composition can reveal about who drives, sustains, or terminates transmission chains in a COVID-19-like pandemic.
